Exosome / Endocytosis / Phagocytosis | Reagent and Kit Selection Guide
Science Note
[Apr. 15, 2026] Previous Science Note
Expansion Microscopy Reveals Native Molecular Landscapes in Cells
|
Expansion microscopy is a useful method that physically expands gel-anchored specimens several-fold, thereby improving the effective spatial resolution of fluorescence imaging and enabling analysis of intracellular nanostructures. At the same time, challenges remain in molecular retention during sample processing and in achieving high-contrast imaging, and various efforts have been made to broaden the scope of ExM. Recently, progress has been reported in two directions. One study introduced Land-ExM, which enabled high-contrast retention of protein and lipid signals together with 3D visualization of organelles and contact sites in combination with immunostaining. Another study developed a fixable and clickable cholesterol probe, enabling nanoscale imaging of cholesterol together with organelle markers. Together, these studies advance ExM as a platform for integrating intracellular molecular distribution with ultrastructural context. |
||||||||||
|
Landscape expansion microscopy reveals interactions between membrane and phase-separated organelles (Journal of Cell Biology, 2026) Highlighted technique: The authors designed land-ExM around NHS-biotin-MA, a trifunctional molecule that reacts with primary amines, covalently anchors labeled components to the hydrogel, and enables post-expansion fluorescent detection through biotin–streptavidin binding. Cellls were fixed and labeled with mCLING for lipids, followed by optional immunostaining, and NHS-biotin-MA was then used to co-anchor proteins and the lipid probe for multichannel ExM imaging. |
||||||||||
|
A Multifunctional Probe for Visualization of the Nanoscale Distribution of Cholesterol in Cells by Expansion Microscopy (Aggregate, 2025) Highlighted technique: Expansion microscopy has advanced nanoscale imaging in cells, but applying it to cholesterol has been difficult because cholesterol is poorly retained during fixation and easily lost during later processing. In this study, the authors expand the scope of ExM by developing a fixable and clickable cholesterol probe, enabling nanoscale visualization of cholesterol together with organelle markers and providing a foundation for extending ExM to other chemically challenging lipid-like small molecules. |
||||||||||
| Related Techniques | ||||||||||
| All-in-one reagents for expansion microscopy | Three-Dimensional Expansion Microscopy Kit (4x 3D-ExM) | |||||||||
| Nuclear staining | DAPI, Hoechst 33342 and Hoechst 33258 | |||||||||
| Related Applications | ||||||||||
Application Note (click to open/close)
|
||||||||||
|
After staining the nuclei and tubulin of RPE1 cells, the cells were expanded using this kit. The samples after expansion were imaged by confocal microscopy with a 60× oil-immersion objective, and 3D imaging was performed. As a result, the detailed morphology of the nuclear structures and cytoskeleton could be observed three-dimensionally at high resolution (Movie 1). Furthermore, two nuclei with different shapes present within the same cell could be compared and their structural differences could be confirmed in three dimensions (Movie 2). Experimental data:Kindly provided by Dr. Aussie Suzuki, University of Wisconsin–Madison.
|
|||||
> Mitochondrial Morphological Changes Induced by Different Drug Treatments
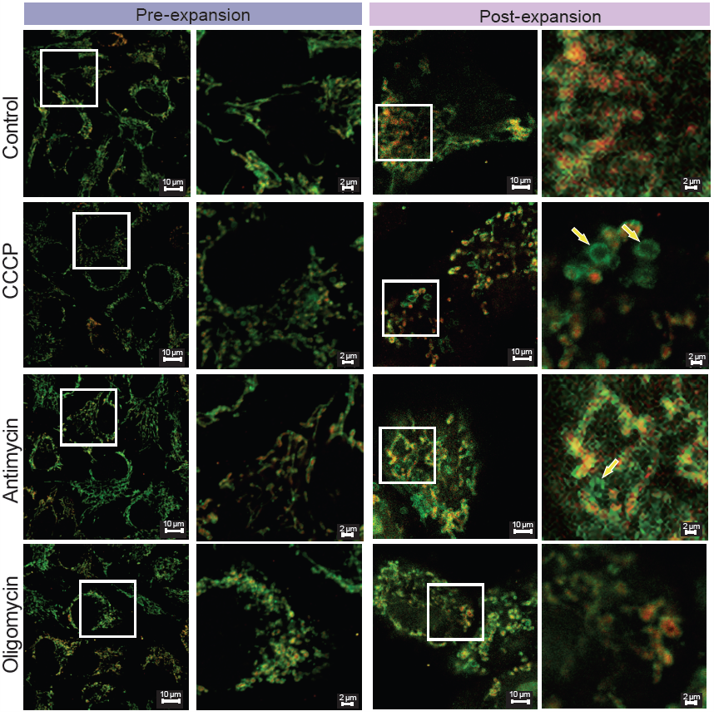 |
<Experimental Conditions> |
What is Plasma Membrane?
The plasma membrane (PM) separates the intracellular environment from the extracellular space and protects the cells from the extracellular environment. The PM plays important roles in selective permeation or signaling between cells etc. Observing the dynamics of the PM is useful in various research field.
Cell Membrane-related Reagents Selectable by Targets

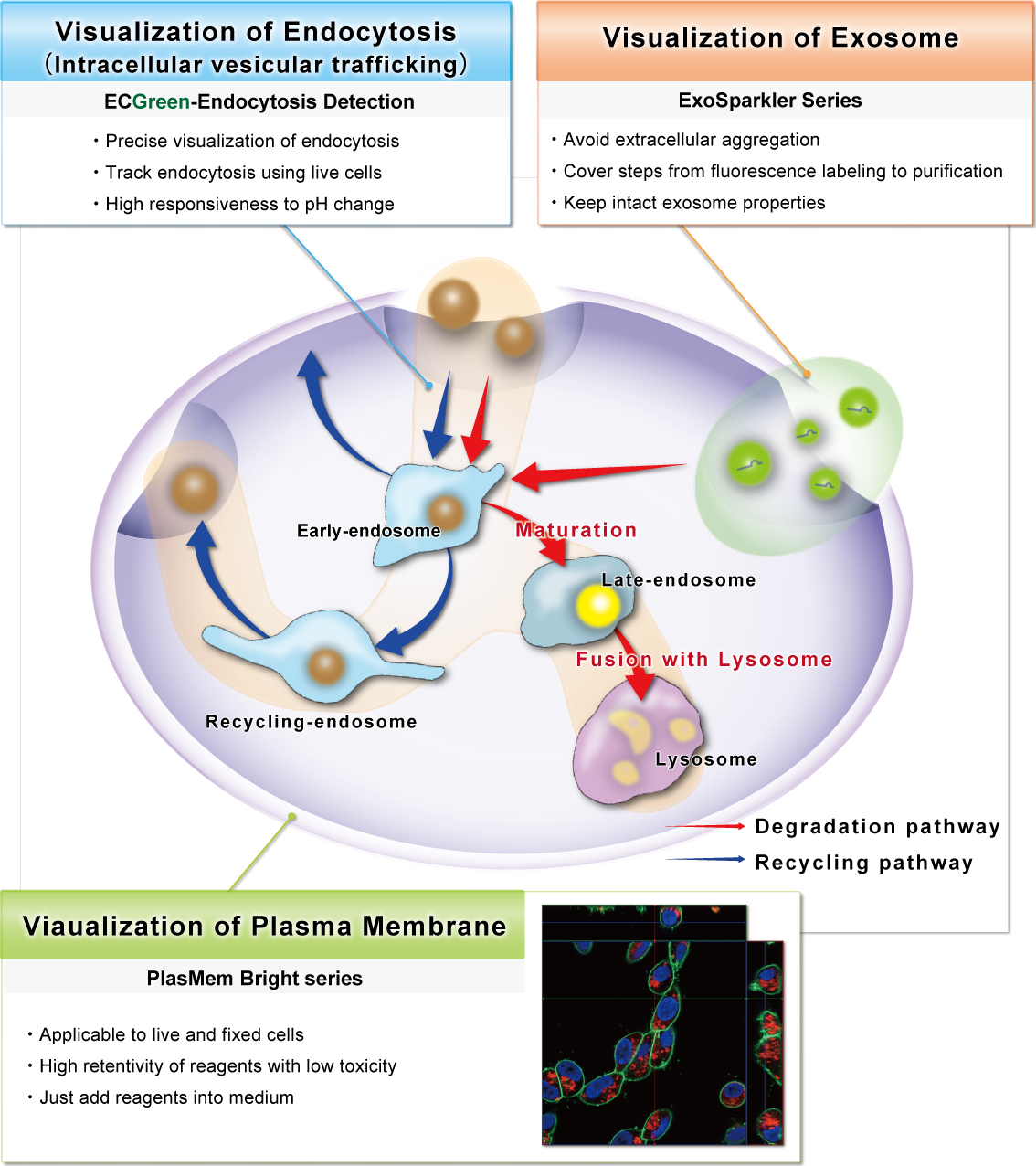
Selesction Guide

~ Features ~
- Applicable to live and fixed cells
- High retentivity of reagents with low toxicity
- Just add reagents into medium
Low toxicity, No washing, and High retentivity
Comparison with other products
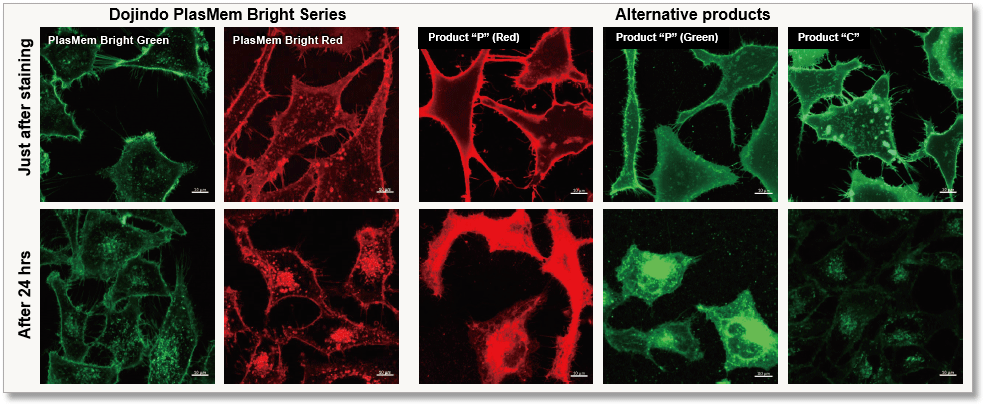
PlasMem Bright Series has low cytotoxicity, and high membrane retention of dyes and can be used in various experiments using live and fixed cells.
Clear visualization of plasma membrane
Observe morphology of neuron (differentiated SH-SY5Y cells) and localization of mitochondria in axon.
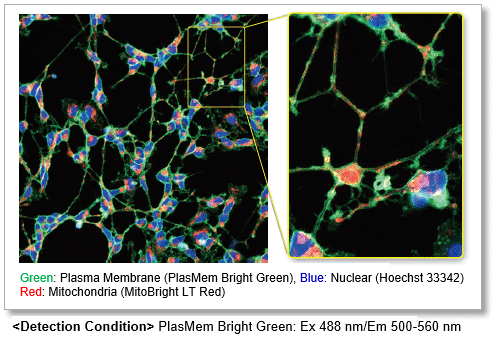
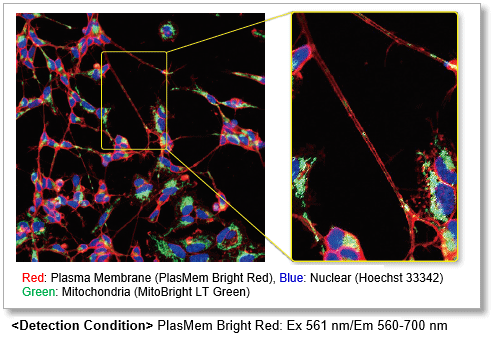
High retentivity on plasma membrane
HeLa cells stained with each plasma membrane staining reagent were incubated for 24 hrs and each the resulting fluorescent image was compared. PlasMem Bright series had higher retentivity in plasma membrane than other products.
| Product | Size | Product Code |
|---|---|---|
| Visualization of Cell membrane | Plasma Membrane Staining Dyes | |
| PlasMem Bright Green | 100 µl x 1 | P504-10 |
| PlasMem Bright Red | 100 µl x 1 | P505-10 |
|
Note: 1 tube (100 µl), 10 assays at 35 mm dish, 10 assays at μ-Slide 8 well |
||
Track endosome

~ Features ~
- Precise visualization of endocytosis
- Track endocytosis using live cells
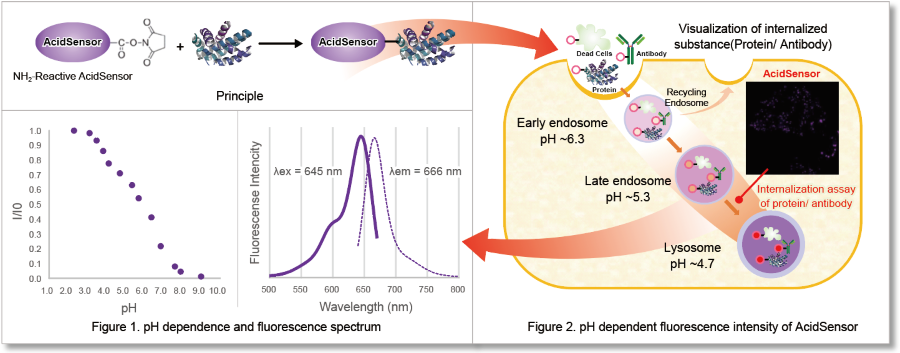
- High responsiveness to pH change
ECGreen is a pH dependent fluorescence dye that localizes to vesicle membrane. The visualization of endocytosis using the ECGreen is a more direct method than fluorescent analogs and allows visualization endocytosis from the stage of early endosomes.
Overall, this results in increased oxidative stress and accelerated cellular damage.
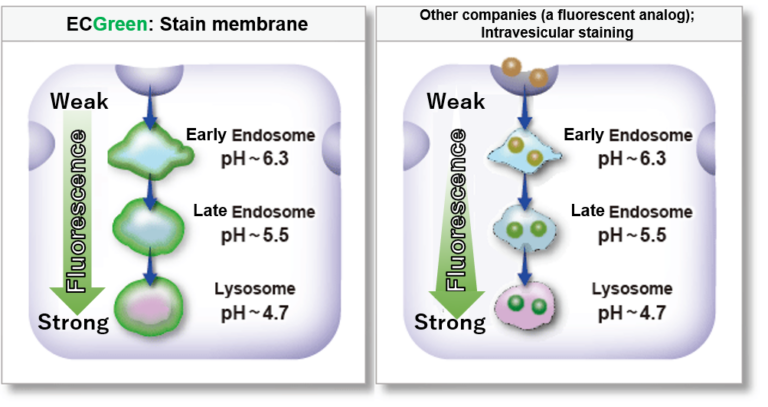
Stain vesicle membrane precisely
Other companies (a fluorescent analog): intravesicular staining
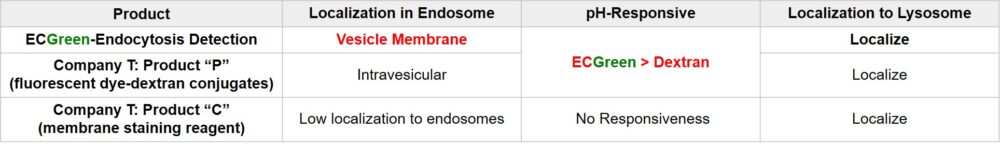
Fluorescent Dye-Dextran Conjugates or membrane staining reagents are used to visualize endocytosis. However, they have limitations in observing dynamics of endosomes in live cells in terms of precision of staining or retentivity of reagent. ECGreen is the reagent that over comes the limitations.
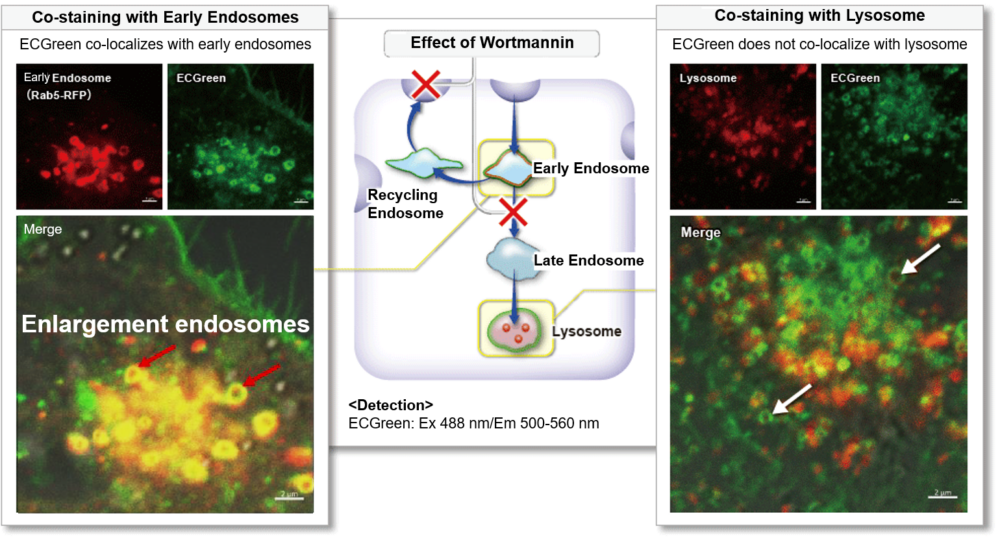
Clear visualization of intracellular vesicular trafficking
It has been known that Wortmannin inhibits the recycling of endosomes or transition to lysosomes and causes enlargement of endosomes. To evaluate these changes caused by Wortmannin, early endosomes were co-stained by ECGreen and Rab5-RFP (marker protein of early endosomes), and lysosomes were co-stained by ECGreen and lysosome staining reagent. In adding Wortmannin, ECGreen was colocalized with enlarged endosomes (Rab5-RFP). On the other hand, ECGreen wasn’t colocalized with lysosomes.
Track Phagocytosis
~ Features ~
- Label with pH-sensitive fluorescent dye
- All-in-one Kit
- Compatible with phagocytosis assay
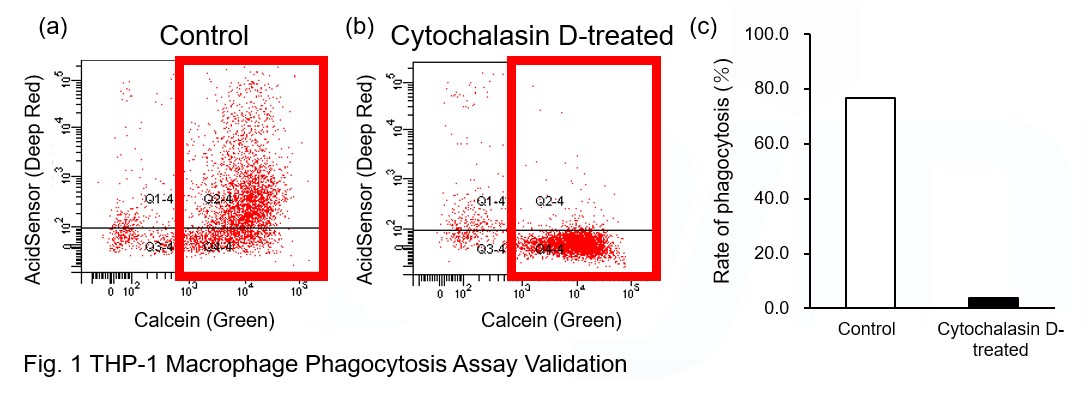
Phagocytosis assay of labeled apoptotic cells in THP-1 cells
AcidSensor-labeled substances are taken up by cells and their fluorescence increases when they reach acidic organelles such as lysosomes. Taking advantage of this property, we evaluate the phagocytic activity of apoptotic cells by co-culturing AcidSensor-labeled apoptotic cells with Calcein-labeled THP-1 macrophages. As a result, Calcein (Green) / AcidSensor (Deep red) double-positive cells, indicating THP-1 macrophages phagocytosing apoptotic cells, were observed by flow cytometry (Fig. 1a). Furthermore, when the phagocytosis of THP-1 macrophages was inhibited by Cytochalasin D, the percentage of double-positive cells decreased (Fig. 1b and 1c), confirming that the assay system can accurately evaluate phagocytosis.
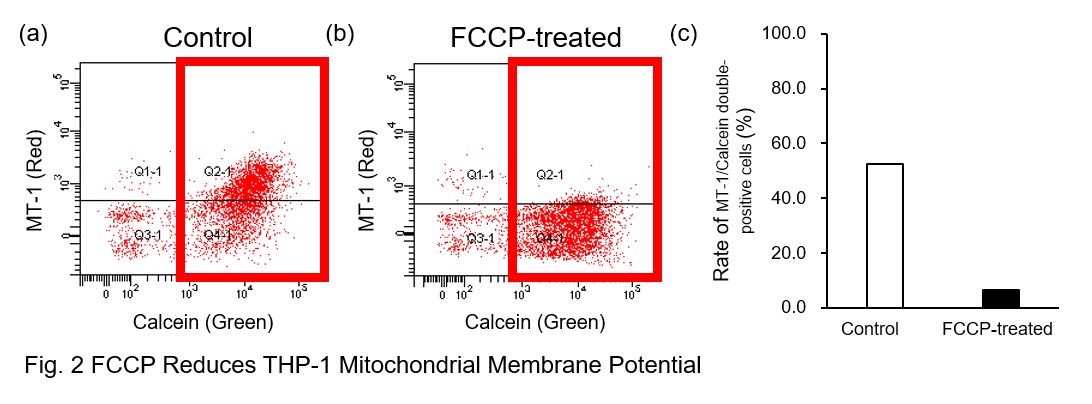
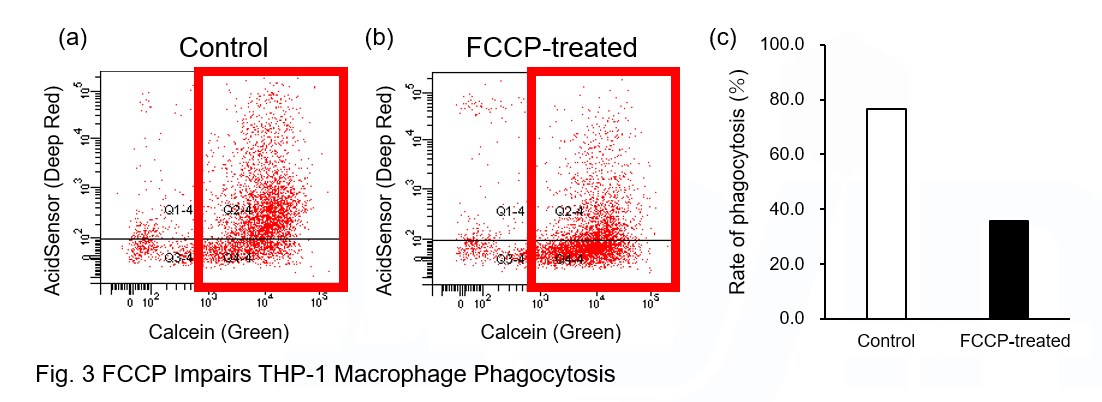
A recent report reveals that inhibition of mitochondrial function induces a switch to glycolysis and reduces phagocytosis in cultured microglia, resident macrophages in the central nervous system*. To replicate this result, phagocytosis assays were performed using mitochondria-inhibited THP-1 macrophages. The results show that FCCP, a potent uncoupler of oxidative phosphorylation in mitochondria, decreases mitochondrial membrane potential (MT-1, Red) of THP-1 macrophages (Fig. 2) and reduces phagocytosis (Fig. 3).
*Lauren H. Fairley, et al., PNAS (2023)
Products in Use
① AcidSensor Labeling Kit – Endocytic Internalization Assay [code: A558]
② -Cellstain- Calcein-AM solution [code: C396]
③ MT-1 MitoMP Detection Kit [code: MT13]
| Product | Size | Product Code |
|---|---|---|
| Visualization of Endocytosis | Endocytosis Detection Dye | |
| ECGreen-Endocytosis Detection | 40 µl x 1 | E296-10 |
|
Note:1 tube (40 µl), 20 assays at 35 mm dish, 20 assays at μ-Slide 8 well |
||
| Product | Size | Product Code |
|---|---|---|
| Visualization of Phagocytosis | Phagocytosis Detection Dye | |
| AcidSensor Labeling Kit | 3 samples | A558-10 |
|
Note:Proteins with a molecular weight above 50,000 can be labeled. |
||
Exosome Membrane/Protein Fluorescence Labeling Kit

~ Features ~
- Not allow extracellular aggregation
- Cover steps from fluorescence labeling to purification
- Little effect on exosome properties
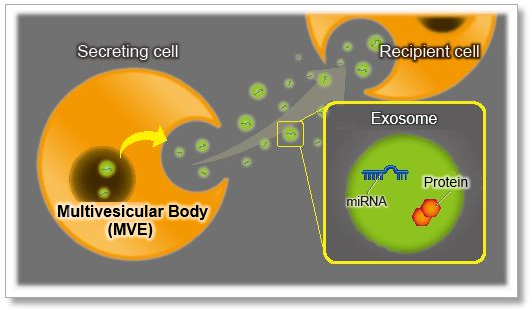
Recent findings suggest that exosomes, a form of extracellular vesicle (EV), contribute to malignant transformation and the metastasis of cancer. Consequently, intercellular communication via exosomes is attracting considerable interest in the scientific community.
To shed light on such communication, labeling techniques based on fluorescent dyes have been used. Fluorescent dyes that label the cellular membrane are commonly used for exosome labeling because the lipid bilayer in exosomes is a good target for labeling.
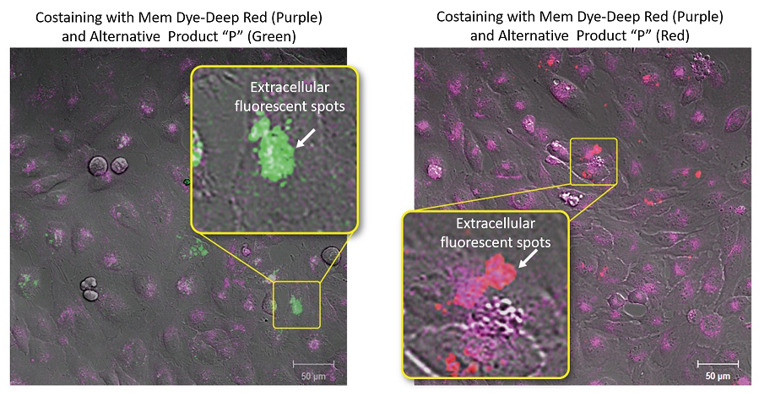
ExoSparkler series does not allow extracellular aggregation
Exosomes stained with ExoSparkler’s Mem Dye-Deep Red or an alternative product (green or red) were added to each well containing HeLa cells. The labeled exosomes taken into HeLa cells were observed by fluorescent microscopy. As a result, extracellular fluorescent spots suspected of dye aggregations were seen in each well containing the exosomes stained with the alternative product (green or red).
Mem Dye-Deep Red (Purple): Ex 640 nm/Em 640-760 nm
Alternative Product “P” (Green): Ex 561 nm/Em 560-620 nm
Alternative Product “P” (Red): Ex 640 nm/Em 650-700 nm
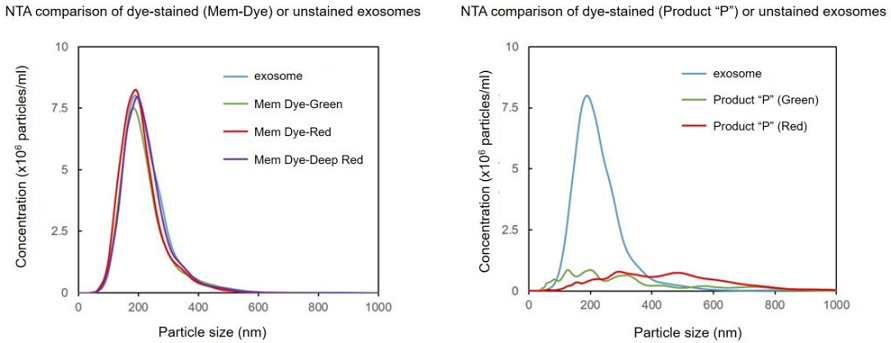
Mem Dye-Deep Red and Product “P” (Green and Red) in aqueous solution were analyzed by NTA (nanoparticle tracking analysis) to investigate the generation of aggregates. No aggregation was observed in the experiments with Mem Dyes, although Product “P” (Green and Red) produced dye-to-dye aggregates (100–500 nm size).
Instrument: LM10-HSBFT 14 (Nanosight)
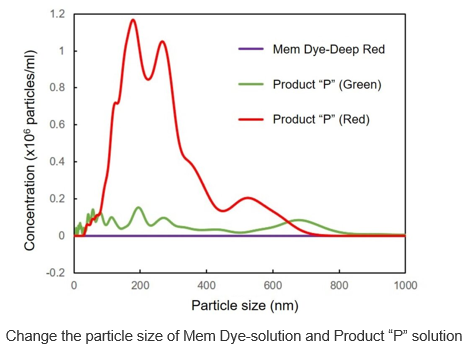
In Mem Dye-Green, Red, the aggregation of the dye was not confirmed as in Mem Dye-Deep Red.
Commonly used exosomal membrane dye can cause dye aggregation, exhibiting fluorescent spots that are not derived from exosomes. These dyes can also change the functional properties of exosomes while increasing the background imaging.1,2
The dyes used in ExoSparkler series (Mem Dye-Green, Red, and Deep Red) do not cause aggregation and have little influence on properties of exosomes, allowing a more accurate observation of exosome dynamics.
1) Mehdi Dehghani et al., “Exosome labeling by lipophilic dye PKH26 results in significant increase in vesicle size”.bioRxiv., 2019, doi:10.1101/532028.
2) Pužar Dominkuš P et al., “PKH26 labeling of extracellular vesicles: Characterization and cellular internalization of contaminating PKH26 nanoparticles.” Biochim Biophys Acta Biomembr., 2018, doi: 10.1016/j.bbamem.2018.03.013.
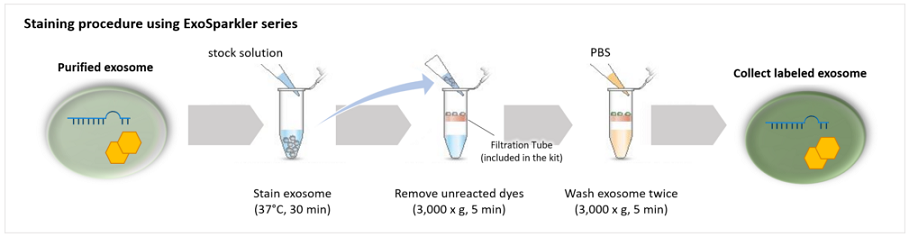
Our ExoSparkler Exosome Membrane Labelling Kits provide everything from fluorescence labeling to purification
ExoSparkler series contains filtration tubes available for the removal of dyes unreacted after fluorescence labeling, as well as an optimized protocol for labeling exosomes. Our ExoSparkler series makes it possible to prepare fluorescence labeling of exosomes using the simple procedure.
Comparison of purification methods (removal of unlabeled dyes)
The filtration tubes used to remove unlabeled dyes in this kit can purify exosomes at a higher recovery rate than gel filtration methods.

For the effectiveness of purification using filtration tubes, please see Q&A.
(The filter is colored in the purification after the labeling, Have unlabeled dyes been removed?)
Mem Dyes have little effect on exosome properties
NTA (nanoparticle tracking analysis) and zeta potential were measured to determine the changes in exosomes of dye-stained with Mem Dye-Deep Red or Product “P” (green or red) or unstained exosomes.
As a result, the Mem-Dye series (green, red, deep red) had little effect on exosome properties.
Effect of the dyes on the particle size of the exosomes
Exosomes were stained with Mem Dye-series (green, red, deep red) and Product “P” (green and red) at a dye concentration of 10 μmol/L in DMSO, the NTA (nanoparticle tracking analysis) of the stained exosomes (as 10 µg protein) was measured.
As a result, Mem Dyes-series did not change number and particle size of the exosomes (bottom left). Conversely, the Product “P” stained exosomes showed the significant changes of particle size and population of the exosomes (bottom right).
Instrument: LM10-HSBFT 14 (Nanosight)
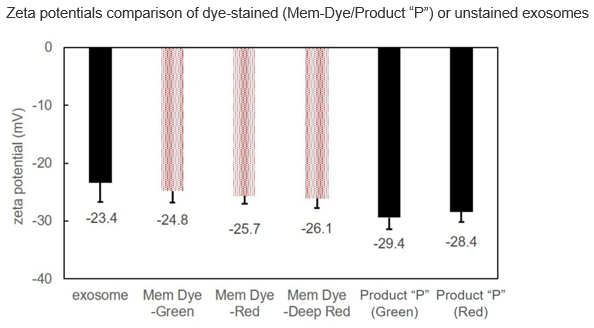
Effect of the dyes on the zeta potentials of the exosomes
Exosomes were stained with Mem Dye-series (green, red, crimson) and Product “P” (green and red) at a dye concentration of 10 μmol/L in DMSO, the zeta potentials of the stained exosomes (as 10 µg protein) was measured.
As a result, product “P”-stained exosomes have lower zeta potential than Mem Dye-stained.
Instrument: Zetasizer Nano ZSP (Malvern Panalytical)
Observethetime-dependent changes in exosome localization
Exosomes purified by ultracentrifugation (10 µg as protein amount) were stained with Mem Dye-Deep Red (Exosome Membrane Fluorescence Labeling Kit) and added to HeLa cells (1.25×104 cells) stained with lysosome staining dye. The fluorescence images were observed after 1 h and 4 h incubation.
As a result, it was confirmed that the fluorescence puncta (purple) of Mem Dye-Deep Red overlapped with the localization of lysosomes (green) over time (white), and that the localization of exosomes changed in a time-dependent manner.
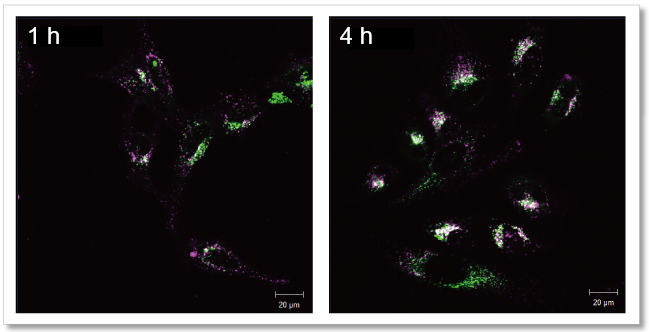
Detection Conditions
Mem Dye-Deep Red (Purple): Ex 640 nm/Em 640-760 nm
Lysosome staining dye: Ex 488 nm/Em 490-540 nm
| Product | Size | Product Code |
|---|---|---|
| Visualization of Exosome | Exosome Membrane Fluorescence Labeling Kits | |
| ExoSparkler Exosome Membrane Labeling Kit-Green | 5 samples | EX01-10 |
| ExoSparkler Exosome Membrane Labeling Kit-Red | 5 samples | EX02-10 |
| ExExoSparkler Exosome Membrane Labeling Kit-Deep Red | 5 samples | EX03-10 |
| Exosome Protein Fluorescence Labeling Kits | ||
| ExoSparkler Exosome Protein Labeling Kit-Green | 5 samples | EX04-10 |
| ExoSparkler Exosome Protein Labeling Kit-Red | 5 samples | EX05-10 |
| ExoSparkler Exosome Protein Labeling Kit-Deep Red | 5 samples | EX06-10 |
|
Note: Protein amount : 1-10μg/ sample, Particle count : 10 to 100 x 108 /samples (As purified exosome using ultracentrifugation) |
||
ExoIsolator Exosome Isolation Kit
~ Features ~
- Easy to use, no technique required
- Recovery rate equivalent to ultracentrifugation
- Filter holder is reusable
Easy to use no technique required
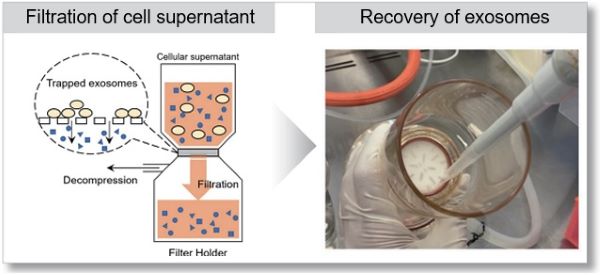
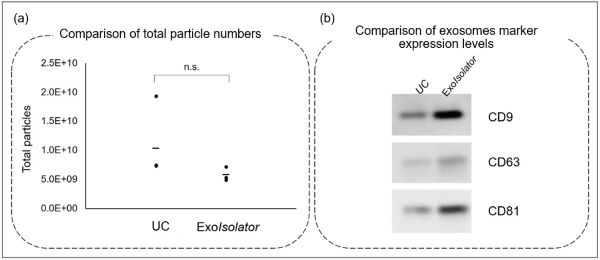
Recovery rate equivalent to ultracentrifugation
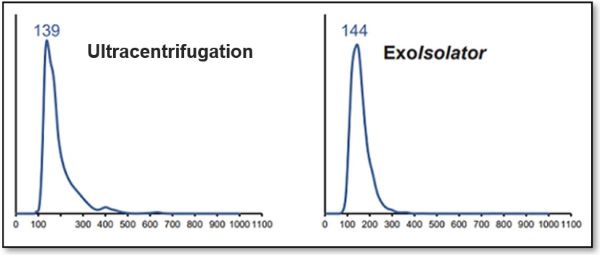
(a) Nanoparticle number and (b) expression level of exosome markers
| Product | Size | Product Code |
|---|---|---|
| ExoIsolator Exosome Isolation Kit | 3 tests | EX10-10 |
| Filter for exosome isolation kit | 10 pieces | EX11-10 |
*ExoIsolator Exosome Isolation Kit contains Filter Holder x 1, Isolation Filter x 3, Tweezers x 1. The Filter Holder can be reused after autoclaving.



















