Science Note
Autophagy Links Tumor Growth and Immune Evasion [Mar. 24, 2026]
|
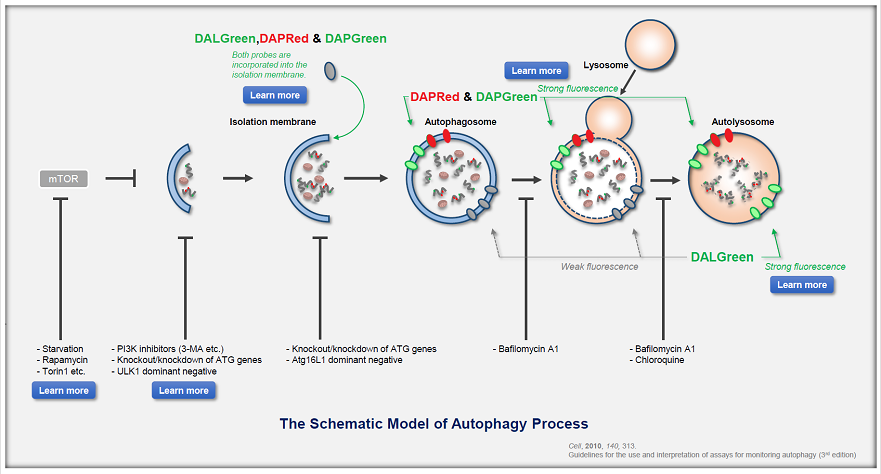
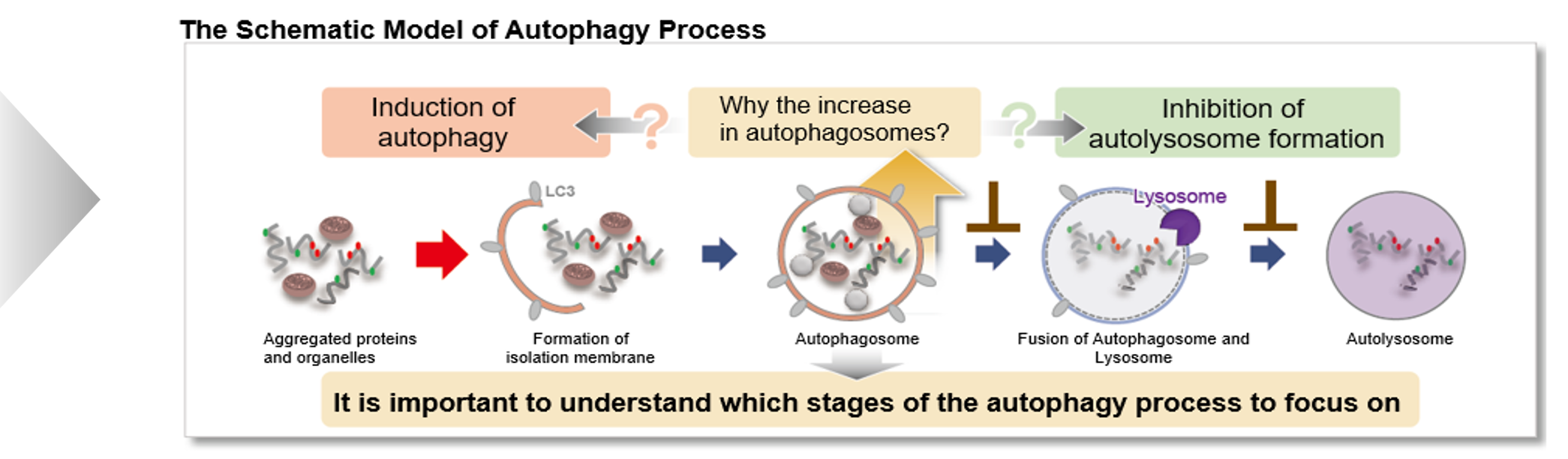
Autophagy supports cellular homeostasis by recycling intracellular components under stress. In cancer, autophagy-related pathways support tumor cell survival through metabolic adaptation and stress tolerance, while also contributing to immune evasion by shaping innate immune signaling and the tumor microenvironment. Recent studies show that glioblastoma stem cells use chaperone-mediated autophagy to degrade the mTORC1 suppressors TSC1/2 and TET3, thereby promoting self-renewal and suppressing innate immune signaling, whereas pancreatic ductal adenocarcinoma depends on the autophagy-initiating kinase ULK1 to maintain autophagic flux, metabolic supply, and expression of immunosuppressive cytokines and chemokines. These studies clarify autophagy-linked mechanisms supporting tumor maintenance and immune suppression. |
||||||||||||||||||
|
1. Targeting chaperone-mediated autophagy inhibits properties of glioblastoma stem cells and restores anti-tumor immunity (Nature Communications, 2025) |
||||||||||||||||||
|
2. ULK1 knockout suppresses pancreatic cancer progression by inhibiting autophagy and enhancing antitumor immunity (Experimental & Molecular Medicine , 2025) |
||||||||||||||||||
|
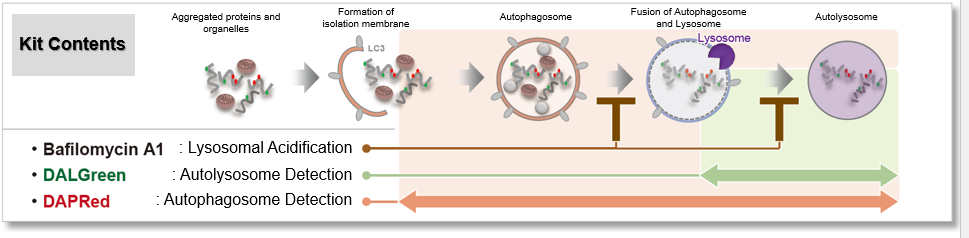
Solutions for Autophagy Experiments |
||||||||||||||||||
Related Techniques (click to open/close)
|
||||||||||||||||||
Application Note I (click to open/close)
|
||||||||||||||||||
|
|
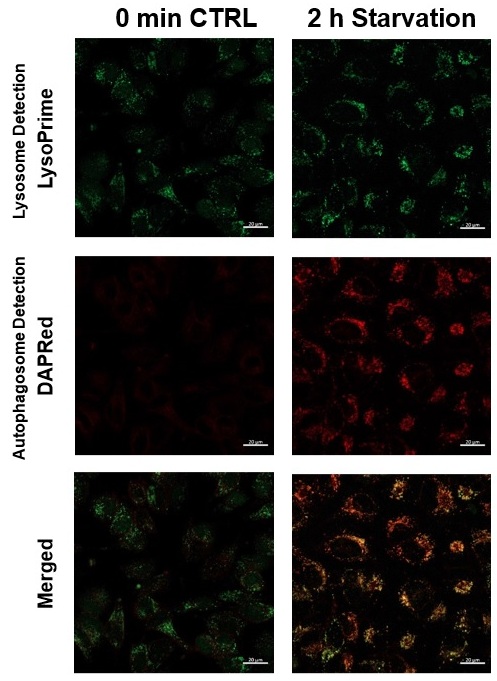
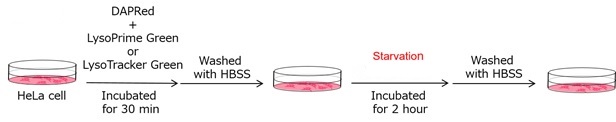
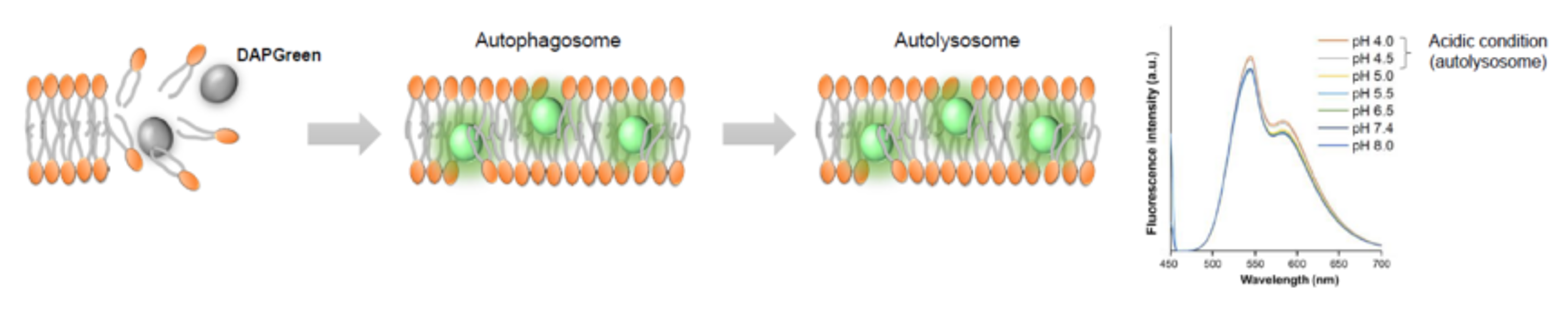
We performed fluorescence imaging under amino acid starvation in HeLa cells stained with the autophagosome detection probe DAPRed (Code: D677) and LysoPrime Green (Code: L261). The fluorescence signal of LysoPrime Green increased, and lysosomal localization was confirmed over time. These results indicate that the co-localization rate with DAPRed fluorescence is sufficient, enabling accurate autophagy analysis. |
Application Note II (click to open/close)
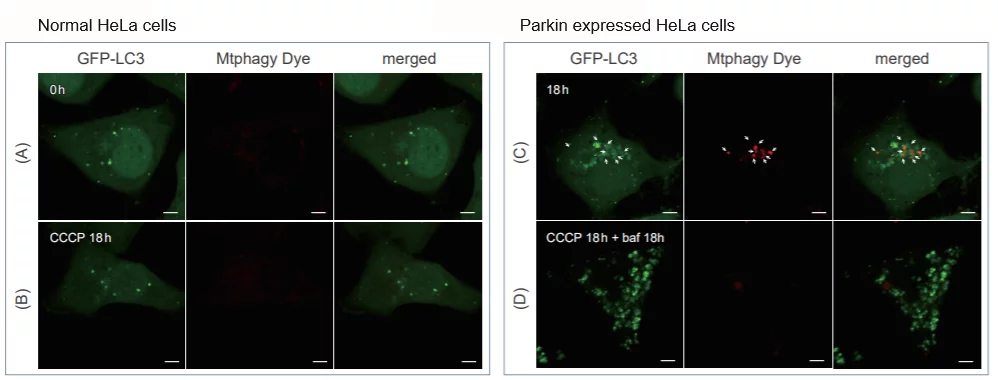
> Induction of Mitophagy in Parkin Expressed HeLa cells
|
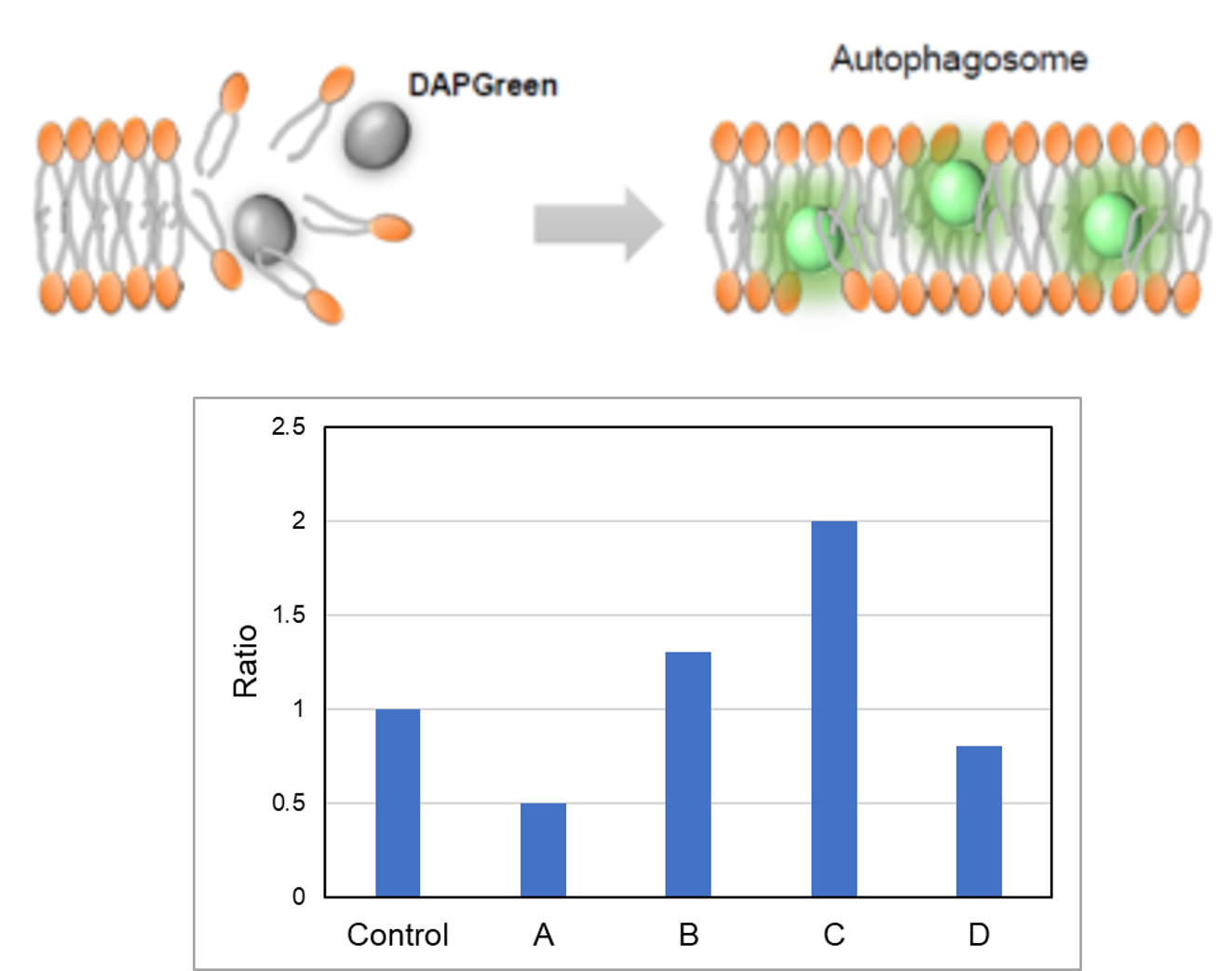
CCCP(carbonyl cyanide m-chlorophenyl hydrazone) has been added to normal and Parkin expressed cells. The strong fluorescence was not observed in normal HeLa cells(A)(B). On the other hand, the trong fluorescence was shown in Parkin expressed cells in 18 hours after additon of CCCP(C). Some of the puncta are co-localized with the autophagy marker(GFP-LC3). In addition, suppressed fluorescence of Mtphagy dye was observed when autophagy inhibitor, bafilomycin was added to Parking expressed cells(D). Because lysosomal pH was increased by the additon of bafilomycin. |